
STRASBOURG - FRANCE
Dear Researchers,
We are pleased to announce the third edition of Open Data at MICCAI 2026.
As medical machine learning continues to advance, access to diverse, representative, and inclusive datasets remains a key bottleneck for addressing global healthcare challenges. The Open Data initiative aims to foster collaboration and innovation by encouraging the sharing of high-quality medical imaging datasets within the MICCAI community.
Building on the success of the first two editions, Open Data at MICCAI 2026 continues to place a strong emphasis on underrepresented populations and diseases. Despite substantial progress in medical imaging research and the growing availability of public datasets, significant gaps persist, particularly for regions such as the Middle East. By showcasing datasets that reflect underrepresented populations and clinical conditions, this initiative seeks to reduce disparities in healthcare research and promote more equitable and inclusive machine learning development.
As in previous years, we are creating the space and community that supports and provides more publicly available, high-quality medical imaging datasets, with a particular focus on underrepresented populations and diseases. Details regarding the submission process and storage guidelines will be shared at a later stage. The initiative remains closely aligned with the principles of FAIR data and Data-centric AI, emphasizing the critical role of well-curated data in building reliable and generalizable models. After all, there are no good models without good data.
New for 2026, thanks to the MICCAI Health Equity Grant, we are pleased to introduce micro-grants for dataset providers. These micro-grants aim to support researchers and institutions in preparing, curating, and sharing high-impact datasets, particularly those representing underrepresented populations and regions. Further details regarding eligibility, application procedures, and grant amounts are provided below.
MICCAI 2026 will take place from September 27th - October 1st, 2026, in Strasbourg. The Open Data event will run in parallel with the main conference. If you are attending MICCAI, you are welcome to visit the Open Data sessions, and no separate registration is required.
Datasets and related works will be selected through a peer-reviewed paper submission process. The MELBA journal will serve as the official journal for submissions from the MICCAI Open Data track.
We look forward to welcoming you to the third Open Data session at MICCAI 2026 and to continuing our collective efforts toward more inclusive and equitable medical AI research.
To maximize the impact of shared medical imaging datasets, we have established a set of clear guidelines for data uploading to ensure accessibility and usability across the research community. By following the instructions below, you can contribute to a well-organized, reusable, and collaborative open-data resource.
Important All data—including imaging and clinical variables—must be anonymized. Any information that could link the data to a patient's identity or medical records must be removed.
If you do not have a dedicated storage space, we recommend using Synapse.
Organize your data logically and consistently:
Use modality-specific data standards wherever possible:
Each dataset must include a README.md file at the root level describing the dataset documentation:
If applicable, provide:
Ideally, at the end of the upload your dataset would have a structure similar to the picture below:
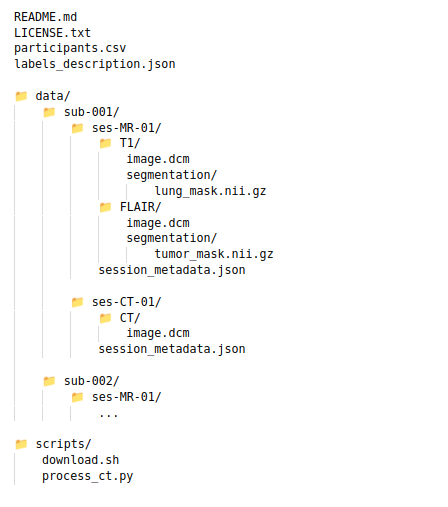
The main focus is on:
Additional topics include:
We welcome submissions of papers presenting novel datasets, particularly those from the Middle East and other underrepresented populations and diseases - including methodologies for the data collection and curation, and innovative approaches for utilizing them in medical imaging research.
CLICK HERE FOR PAPER SUBMISSION SITE
We invite authors to carefully review our updated author guidelines document:
PLEASE CLICK HERE TO VIEW OPEN DATA MICCAI 2026 AUTHOR GUIDELINES
New this year: to improve our review process, we now require the authors to provide the link to their repository upon submission. If the dataset is going to be shared with restricted access, the authors are invited to provide a private link or create a reviewer account for the Open Data 2026 reviewers to be able to access and review the repository.
For more information on licensing and ethics, please read the information in the guide below:
Authors are invited to share their papers as pre-prints, including during the submission phase.
This is a single-blind review process. For general guidelines on "What Makes a Good Review" please refer to the corresponding section in the MICCAI reviewer guidelines.
For this track, pay special attention on the Submission guidelines listed above - scope and criteria, summarized below (in order of priority):
We are pleased to announce Open Data Micro-Grants as part of the Health Equity Award.
These grants support the creation of original, publicly available open datasets in medical imaging and clinical AI, with a focus on health equity, diversity, and under-represented populations.
Travel costs are not eligible.
Applications are closed
PLEASE CLICK HERE TO VIEW OPEN DATA MICRO-GRANTS 2026 RESULTS
The Microsoft CMT service was used for managing the peer-reviewing process for this conference. This service was provided for free by Microsoft and they bore all expenses, including costs for Azure cloud services as well as for software development and support.